Reasoning Engine
An AI-Assisted Differential Diagnosis System for Ophthalmology
Presentation at VietAI Summit 2018.
Demo video.
Background
Late 2019, after successfully building several image analysis models (both classification, segmentation, and object detection to detect lesions in retinal color fundus photographs) using Deep Convolutional Neural Networks and establishing a pipeline for collecting and cleaning data, it dawned on us that we were nowhere close to our ultimate goal, which is a comprehensive diagnosis system, since such a system requires more than just imaging data.
If we had a lot of structured clinical data, it would be quite straightforward since we could use statistical methods to incorporate clinical knowledge into a ML model, and we could even train on clinical data and imaging data together.
But such data was not available where we work, and naturally in many places in developing countries as well, since EHR data was not intentionally built for this purpose.
This forced us to explore a different technology, one that allows us to mimic doctors’ clinical reasoning without having to rely on a large amount of historical data. Knowledge Graphs seemed to us a viable path.
Motivation
Several recurring pain points in daily clinical practice at Cao Thang Eye Hospital motivated this project:
- Repetitive patient interviewing. Doctors spend a significant portion of each consultation asking the same intake questions, regardless of the patient’s presenting condition.
- Diagnostic errors by junior clinicians. Less experienced doctors, lacking sufficient clinical exposure, are prone to missing diagnoses or over-relying on a narrow differential.
- Incomplete EMR data entry. Nursing staff frequently omit relevant data when filling Electronic Medical Records (EMR), leaving gaps that make follow-up harder and reduce the quality of retrospective analysis.
- Irrelevant medical forms. Traditional intake forms are static and ask every patient the same questions, wasting time and missing condition-specific detail.
- Limited access in remote areas. Patients outside major cities have little access to specialist ophthalmology care, making remote screening and triage a critical unmet need.
The goal is a system that addresses all five: it auto-interviews patients adaptively, provides a ranked differential diagnosis with reasoning, reduces the cognitive load on junior clinicians, guides nurses toward complete data entry, and ultimately serves as the AI core for remote satellite clinics.
System Overview
The system takes all available patient data as input and returns a ranked list of probable diagnoses together with an explanation for each result. The inputs span four categories:
- Patient-reported data: personal information (age, gender, race), medical history, and chief complaints.
- Physician observations: clinical signs elicited during examination.
- Device parameters: IOP, visual acuity, and other quantitative measurements.
- Images: color fundus photographs (FO), optical coherence tomography (OCT), and fluorescein angiography (FA), processed by an image reading tool.
The two core components are an image reading tool that extracts abnormal findings from ophthalmic images, and a reasoning engine grounded in a structured knowledge base (KB) that captures relationships between symptoms, signs, risk factors, and diseases. The reasoning engine combines all inputs against the KB to score candidate diseases and surface an interpretable differential.
Knowledge Base
Design
The knowledge base uses a deliberately simple schema with two entity types, Disease and Observation, connected by typed relations. Observations are divided into three subtypes:
- Risk Factors — background conditions that predispose a patient to disease (e.g., diabetes, hypertension).
- Symptoms — subjective experiences reported by the patient (e.g., blurred vision, eye pain).
- Signs — objective findings that only a clinician or imaging tool can detect (e.g., drusen on a fundus photograph, elevated IOP).
A disease can itself serve as an observation of another disease, reflecting the causal chains and comorbidities common in ophthalmology. Despite the simplicity of the entity model, the resulting graph is highly connected: the current KB contains 556 diseases, 1,233 observations, and 2,306 relations. A snapshot of the knowledge graph is shown in Figure 1.
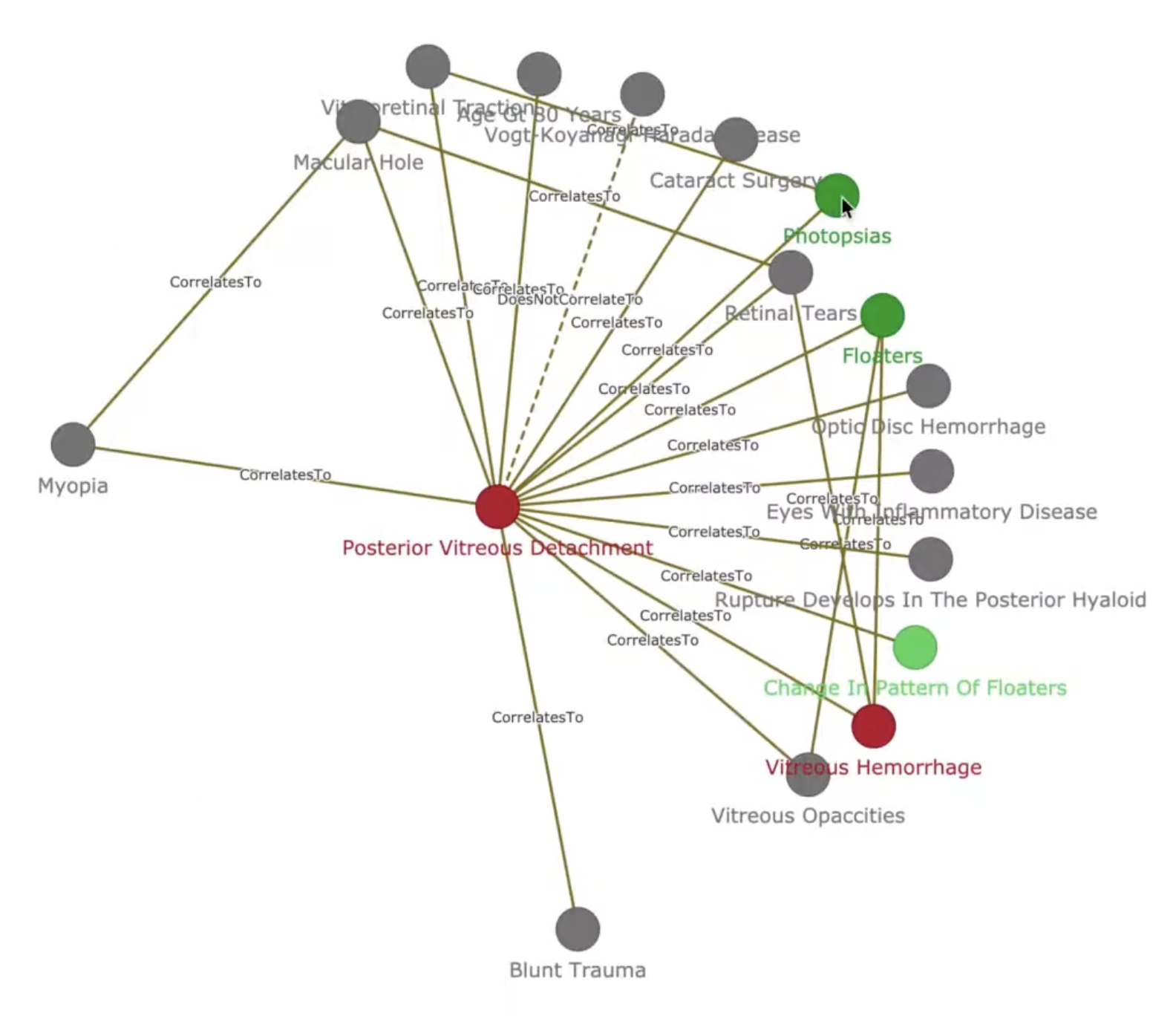
Construction
The initial KB was built from 42 annotated disease articles covering the 50 most prevalent conditions in the CTEH Electronic Medical Records, selected by frequency of occurrence in the hospital’s patient population. The primary content source is the American Academy of Ophthalmology’s EyeWiki. The 42 annotated diseases are listed in Table 1.
| # | Disease | # | Disease |
|---|---|---|---|
| 1 | Ocular Neuropathic Pain | 22 | Cataract |
| 2 | Retinitis Pigmentosa | 23 | Myopia |
| 3 | Presbyopia | 24 | Pigment Epithelial Detachment |
| 4 | Posterior Vitreous Detachment | 25 | Branch Retinal Artery Occlusion |
| 5 | Fungal Keratitis | 26 | Juvenile Open-Angle Glaucoma |
| 6 | Cystoid Macular Edema | 27 | Giant Retinal Tears |
| 7 | Vitreomacular Traction Syndrome | 28 | Retinal Vasculitis |
| 8 | Optic Atrophy | 29 | Peripheral Retinal Degenerations |
| 9 | Neovascular Glaucoma | 30 | Chalazion |
| 10 | Scleritis | 31 | Conjunctivitis |
| 11 | Primary Angle-Closure Glaucoma | 32 | Blepharoptosis |
| 12 | Branch Retinal Vein Occlusion | 33 | Diabetic Retinopathy |
| 13 | Acute Anterior Uveitis | 34 | Primary Open-Angle Glaucoma |
| 14 | Hyperopia | 35 | Retinal Artery Occlusion |
| 15 | Macular Hole | 36 | Bacterial Conjunctivitis |
| 16 | Vitreous Hemorrhage | 37 | Central Retinal Vein Occlusion |
| 17 | Diabetic Macular Edema | 38 | Optic Disc Hemorrhage |
| 18 | Amblyopia | 39 | Age-Related Macular Degeneration |
| 19 | Abducens Nerve Palsy | 40 | Vogt-Koyanagi-Harada Disease |
| 20 | Uveitic Glaucoma | 41 | Toxocariasis |
| 21 | Polypoidal Choroidal Vasculopathy | 42 | Blepharitis |
One limitation of extracting knowledge purely from medical literature is that published texts rarely quantify the strength of association between a finding and a disease. To address this, we are building an annotation interface that allows ophthalmologists to directly encode their clinical experience into the KB, supplying correlation weights that textbooks alone cannot provide.
Milestones
The project is structured around three progressive milestones:
Milestone 1 — Knowledge Base and Disease Ranking. Construct the knowledge graph and implement a scoring function that computes the probability of each candidate disease given an observed set of findings. This is the foundation for all downstream reasoning.
Milestone 2 — Adaptive Auto-Interview System. Replace the static intake form with a dynamic question-generation system. Given a patient’s current set of answers, the system selects the most informative follow-up from one of three action types:
- Elaboration (OLDCARTS): Ask for more detail about a reported complaint, covering onset, location, duration, character, aggravating and relieving factors, timing, and severity.
- New symptom elicitation: Proactively ask about symptoms the patient has not mentioned but that are diagnostically relevant given the current evidence.
- Test or imaging referral: When the most informative next observation requires a clinical measurement or image, prompt the patient to undergo the appropriate test (e.g., fundus photography, OCT) and pipe the image reading tool’s output back into the reasoning loop.
Milestone 3 — Multimodal Integration. Fuse the structured KB with the image reading tool to improve disease prediction accuracy. The KB provides a semantic scaffold for interpreting imaging findings, while the imaging tool surfaces visual signs that enrich the graph traversal.
Open Challenges
Several non-trivial problems remain before the system is clinically deployable:
- Knowledge base completeness and quality. The current KB covers 42 of the 50 target diseases. Expanding coverage, validating extracted relations against clinical ground truth, and establishing a principled review process are ongoing tasks.
- Natural language understanding. The auto-interview system requires robust NLU to parse free-text patient responses, handle negation (“no pain”), resolve synonyms, and generate fluent follow-up questions in both Vietnamese and English.
- Reasoning algorithms. Scoring disease probabilities over a graph with hundreds of nodes and typed relations is a non-trivial inference problem. Approaches range from rule-based propagation to probabilistic graphical models and graph neural networks, and the right choice depends on data availability and interpretability requirements.
- Evaluation. Standard classification metrics are insufficient for a differential diagnosis system. Appropriate evaluation must account for the rank ordering of the differential, the clinical cost of false negatives, and the system’s ability to produce explanations a clinician can audit.